-Search query
-Search result
Showing all 42 items for (author: laughlin & tg)

EMDB-25201: 
H. neapolitanus carboxysomal rubisco/CsoSCA-peptide (1-50)complex
Method: single particle / : Blikstad C, Dugan E

EMDB-25228: 
H. neapolitanus carboxysomal rubisco/CsoSCA-peptide (1-50)complex
Method: single particle / : Blikstad C, Dugan E

EMDB-27973: 
Erwinia amylovora 70S Ribosome
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-27993: 
Erwinia amlyavora 50S ribosome
Method: subtomogram averaging / : Prichard A, Lee J, Laughlin TG, Lee A, Thomas K, Sy A, Spencer T, Asavavimol A, Cafferata A, Cameron M, Chiu N, Davydov D, Desai I, Diaz G, Guereca M, Hearst K, Huang L, Jacobs E, Johnson A, Kahn S, Koch R, Martinez A, McDonald-Uyeno A, Norquist M, Pau T, Prasad G, Saam K, Sandu M, Sarabia A, Schumaker S, Sonin A, Zhao A, Corbett A, Pogliano K, Meyer J, Grose J, Villa E, Dutton R, Pogliano J

EMDB-28003: 
Erwinia phage vB_EamM_RAY (RAY) Capsid Vertex
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28004: 
Erwinia phage vB_EamM_RAY (RAY) Capsid Collar
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28005: 
Erwinia phage vB_EamM_RAY (RAY) Tail Sheath
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28006: 
Erwinia phage vB_EamM_RAY (RAY) Baseplate
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28007: 
Erwinia phage vB_EamM_RAY (RAY) Chimallin
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28008: 
Erwinia phage vB_EamM_RAY (RAY) Putative PhuZ Filament
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28009: 
Pseudomonas chlororaphis phage 201phi2-1 PhuZ Filament
Method: subtomogram averaging / : Laughlin TG, Villa E

EMDB-28010: 
Pseudomonas phage phiKZ PhuZ Filament
Method: subtomogram averaging / : Laughlin TG, Villa E
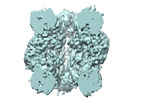
EMDB-27654: 
A subtomogram average of H. neapolitanus Rubisco within alpha-carboxysomes
Method: subtomogram averaging / : Metskas LA, Blikstad C, Laughlin T, Savage DF, Jensen GJ

EMDB-25183: 
P. chlororaphis 70S ribosome in situ subtomogram average
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25220: 
In situ subtomogram average of the 201phi2-1 phage nucleus major shell protein, chimallin (concave class)
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25221: 
In situ consensus subtomogram average of the 201phi2-1 chimallin
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25222: 
In situ subtomogram average of 201phi2-1 phage nucleus major shell protein, chimallin (intermediate/flat class)
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25223: 
In situ subtomogram average of the 201phi2-1 phage nucleus major shell protein, chimallin (convex class)
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25229: 
In situ subtomogram average of the Goslar major phage nucleus shell protein, chimallin (consensus class)
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25262: 
In situ subtomogram average of Goslar phage nucleus major shell protein, chimallin (concave class)
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25358: 
In situ subtomogram average of the major Goslar phage nucleus shell protein, chimallin (convex class)
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25359: 
In situ subtomogram average of the APEC2248 70S ribosome
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25360: 
In situ subtomogram average of the APEC2248 50S ribosome
Method: subtomogram averaging / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25390: 
201Phi2-1 Chimallin Cubic (O, 24mer) assembly
Method: single particle / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25391: 
201phi2-1 Chimallin localized tetramer reconstruction
Method: single particle / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25392: 
201phi2-1 Chimallin C1 localized reconstruction
Method: single particle / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25393: 
201phi2-1 chimallin rectangular (D4,40mer) assembly
Method: single particle / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25394: 
Goslar chimallin cubic (O, 24mer) assembly
Method: single particle / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25395: 
Goslar chimallin C4 tetramer localized reconstruction
Method: single particle / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-25396: 
Goslar chimallin C1 localized reconstruction
Method: single particle / : Laughlin TG, Deep A, Prichard AM, Seitz C, Gu Y, Enustun E, Suslov S, Khanna K, Birkholz EA, Amaro RE, Pogliano J, Corbett KD, Villa E

EMDB-22518: 
SpCas9 delta4CE Ternary Complex
Method: single particle / : Sham A, Laughlin TG, Savage DF

EMDB-20207: 
Synthetic beta-carboxysome shell (T=4)
Method: single particle / : Sutter M, Laughlin TG, Sloan NB, Serwas D, Davies KM, Kerfeld CA

EMDB-20208: 
Synthetic beta-carboxysome shell (T=3)
Method: single particle / : Sutter M, Laughlin TG, Sloan NB, Serwas D, Davies KM, Kerfeld CA

EMDB-20209: 
Synthetic beta-carboxysome shell (prolate full)
Method: single particle / : Sutter M, Laughlin TG, Sloan NB, Serwas D, Davies KM, Kerfeld CA

EMDB-20210: 
Synthetic beta-carboxysome shell (T=4)
Method: single particle / : Sutter M, Laughlin TG, Sloan NB, Serwas D, Davies KM, Kerfeld CA

EMDB-0415: 
T.elongatus NDH (data-set 1)
Method: single particle / : Laughlin TG, Bayne A, Trempe JF, Savage DF, Davies KM

EMDB-0416: 
T.elongatus NDH Peripheral Arm Focus Map(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0417: 
T.elongatus NDH Peripheral Arm Focus Map with NdhS(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0418: 
T.elongatus NDH Peripheral Arm Focus Map without NdhS(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0419: 
T.elongatus NDH Peripheral Arm Focus Map with X-cofactor(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0420: 
T.elongatus NDH Peripheral Arm Focus Map without X-cofactor(data-set 1)
Method: single particle / : Laughlin TG, Davies KM
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
